Quantitative analysis of biochemical processes in living cells at a single-molecule level: a case of olaparib–PARP1 (DNA repair protein) interactions
A. Karpińska, M. Pilz, J. Buczkowska, P. J. Żuk, K. Kucharska, G. Magiera, K. Kwapiszewska, and R. Hołyst
Analyst, 2021,146, 7131-7143
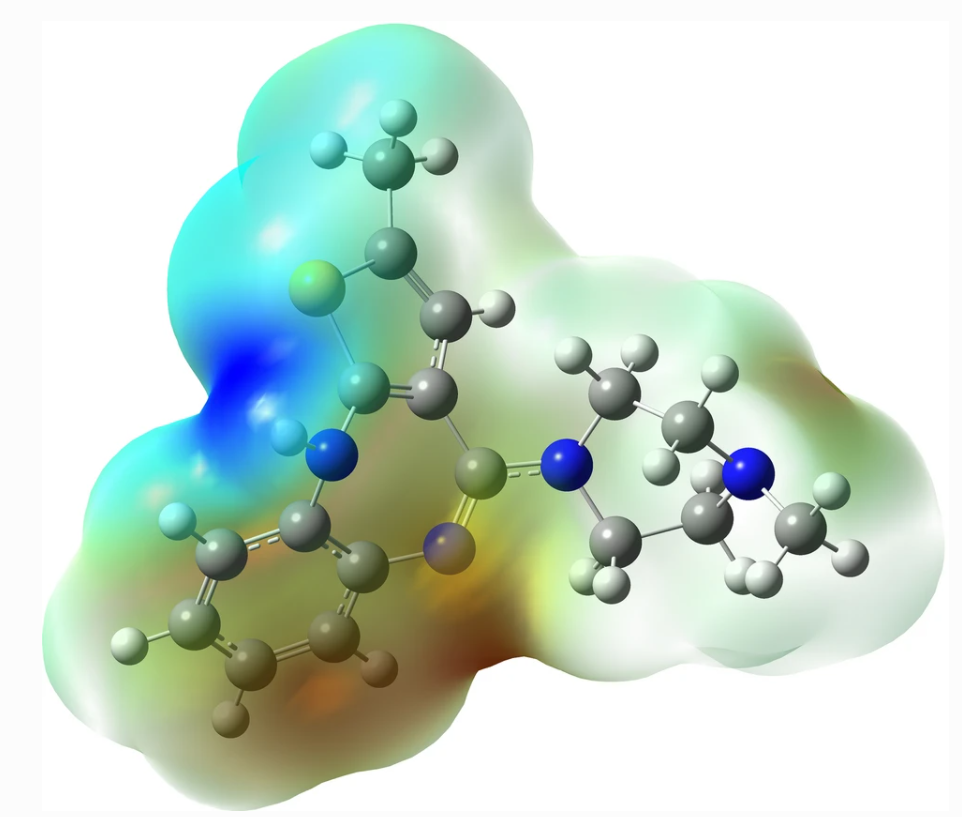
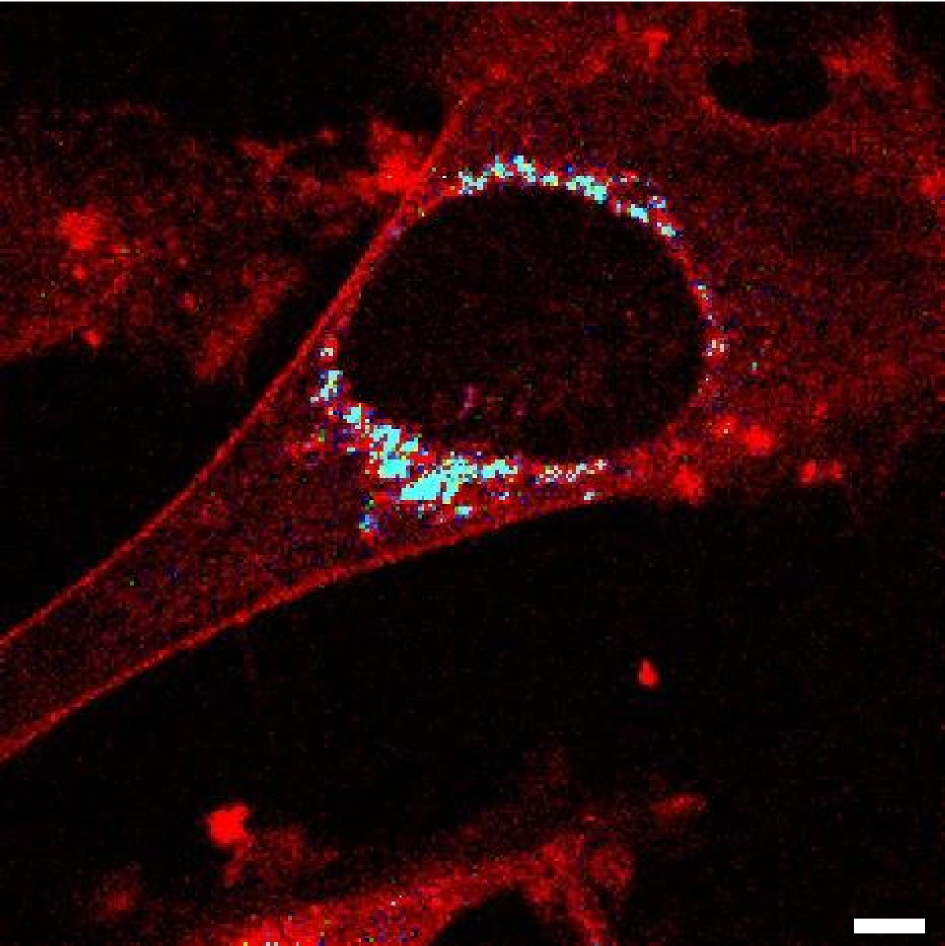
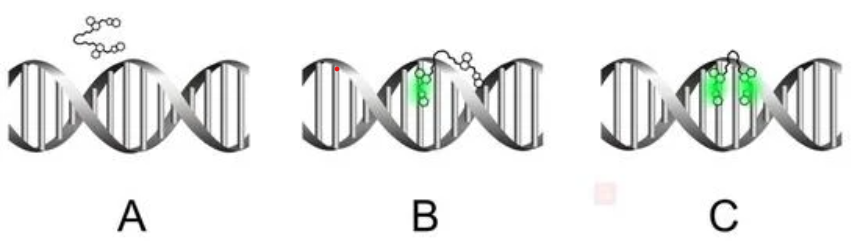
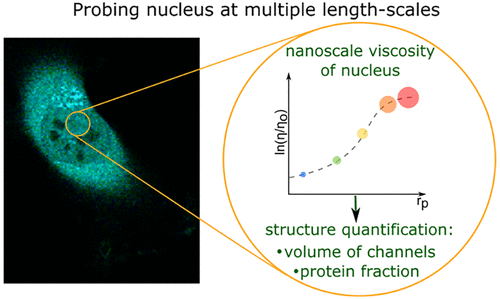
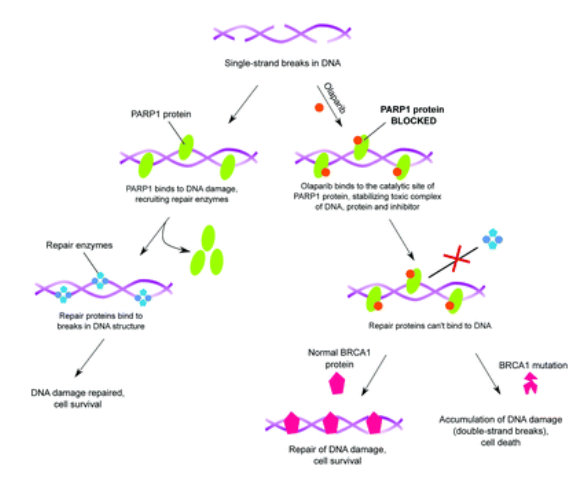
Quantitative description of biochemical processes inside living cells and at single-molecule levels remains a challenge at the forefront of modern instrumentation and spectroscopy. This paper demonstrates such single-cell, single-molecule analyses performed to study the mechanism of action of olaparib – an up-to-date, FDA-approved drug for germline-BRCA mutated metastatic breast cancer. We characterized complexes formed with PARPi-FL – fluorescent analog of olaparib in vitro and in cancer cells using the advanced fluorescent-based method: Fluorescence Correlation Spectroscopy (FCS) combined with a length-scale dependent cytoplasmic/nucleoplasmic viscosity model. We determined in vitro olaparib–PARP1 equilibrium constant (6.06 × 108 mol L−1). In the cell nucleus, we distinguished three states of olaparib: freely diffusing drug (24%), olaparib–PARP1 complex (50%), and olaparib–PARP1–RNA complex (26%). We show olaparib accumulation in 3D spheroids, where intracellular concentration is twofold higher than in 2D cells. Moreover, olaparib concentration was tenfold higher (506 nmol L−1vs. 57 nmol L−1) in cervical cancer (BRCA1 high abundance) than in breast cancer cells (BRCA1 low abundance) but with a lower toxic effect. Thus we confirmed that the amount of BRCA1 protein in the cells is a better predictor of the therapeutic effect of olaparib than its penetration into cancer tissue. Our single-molecule and single-cell approach give a new perspective of drug action in living cells. FCS provides a detailed in vivo insight, valuable in drug development and targeting.