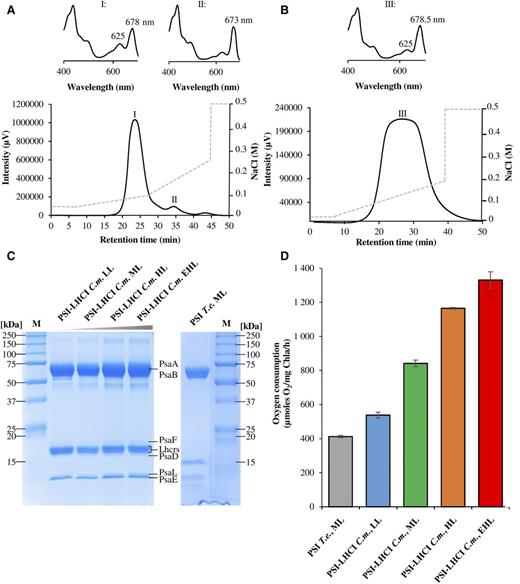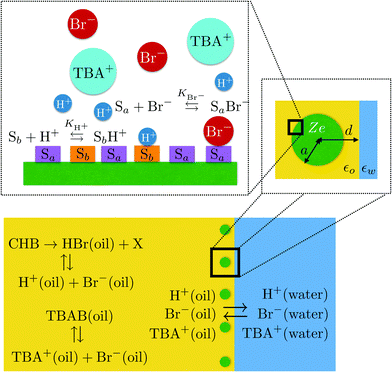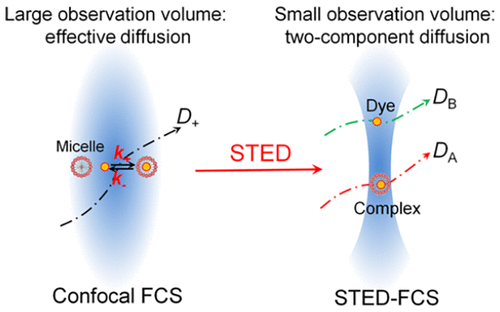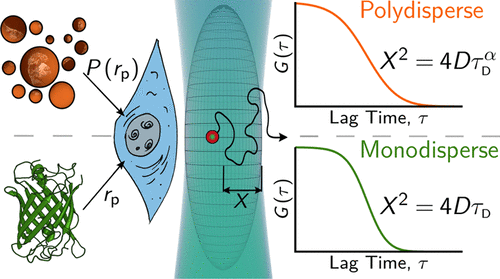
Orientation of photosystem I on graphene through cytochrome c553 leads to improvement in photocurrent generation
M. Kiliszek, E. Harputlu, M. Szalkowski, D. Kowalska, C. G. Unlu, P. Haniewicz, M. Abram, K. Wiwatowski, J. Niedziółka-Jönsson, S. Maćkowski, K. Ocakoglu and J. Kargul
J. Mater. Chem. A, 2018,6, 18615-18626
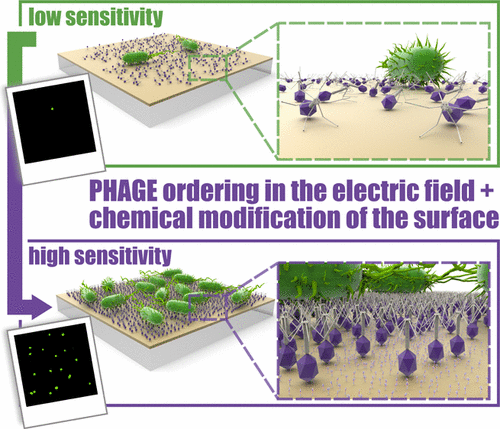
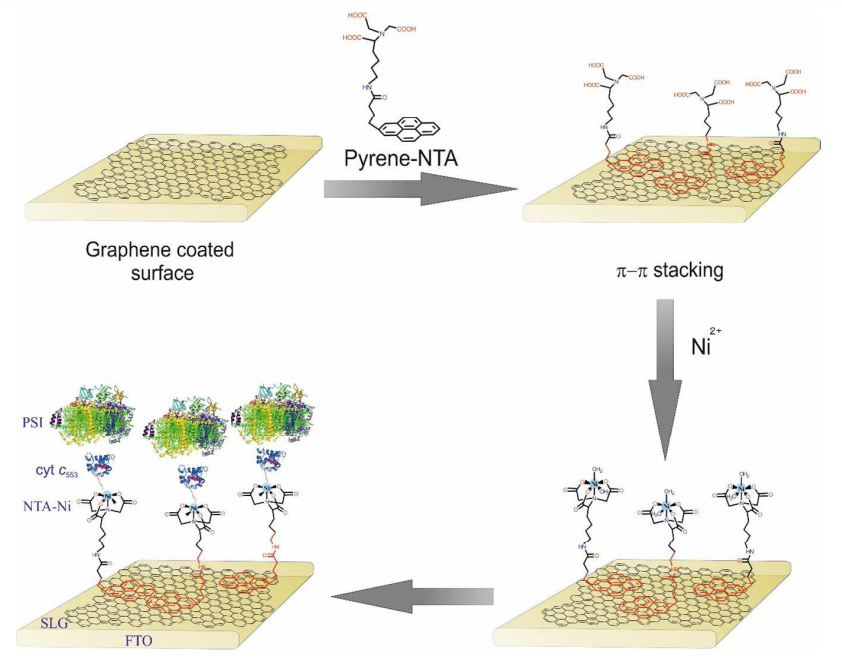
We report the fabrication of an oriented bioelectrode of photosystem I (PSI) on single-layer graphene (SLG). This bioelectrode demonstrates improved photocurrent generation, which can be directly attributed to the molecular conductive interface formed by cytochrome c553 (cyt c553) promoting the uniform orientation of PSI with its donor side towards the electrode. The conductive interface between PSI-cyt c553 and SLG is facilitated by a monolayer composed of π–π-stacked pyrene functionalized with the Ni-NTA moiety, which binds the His6-tagged cyt c553. The surface uniformity of the PSI protein orientation in the electrode structure is evidenced by cross-sectional scanning electron microscopy and fluorescence microscopy, with the latter also proving the efficient electronic coupling between majority of the PSI complexes and graphene. With the uniform organization of the biological photoactive layer, photocurrents are generated at the open circuit potential, which can be further increased when a negative potential is applied. Indeed, at the highest applied negative potential (−0.3 V), over 5-fold increase in the cathodic photocurrent for the PSI complexes conjugated via cyt c553 to the SLG substrate is observed compared with that obtained for the randomly oriented structure where PSI is physisorbed on graphene. These results indicate the key role of a strictly defined orientation of photoactive proteins on electrodes for proper electron transfer and substantial improvement in photocurrent generation in the present or similar bioelectrode architectures.